Multiple high-quality genomes assembled from 24 wild and 20 cultivated potato varieties

A team of researchers affiliated with multiple institutions in China, Europe and the Americas has assembled 44 high-quality genomes from 24 wild and 20 cultivated potato varieties.
In their paper published in the journal Nature, the group describes their sequencing of potato varieties and subsequent analysis. Juanita Gutiérrez-Valencia and Tanja Slotte, with Stockholm University have published a News & Views piece in the same journal issue outlining the work and explaining why such studies have become more important in recent years.
As Gutiérrez-Valencia and Slotte note, potatoes are the third-most consumed crop in the world, behind only wheat and rice. That makes them "the world's most important non-cereal food crop," according to the researchers.
It is therefore critical to learn more about them as global warming threatens to change the ways and possibly the places where potatoes can be grown. To that end, the researchers conducted an extensive genomic study of 24 wild varieties and 20 landraces (cultivated potatoes).
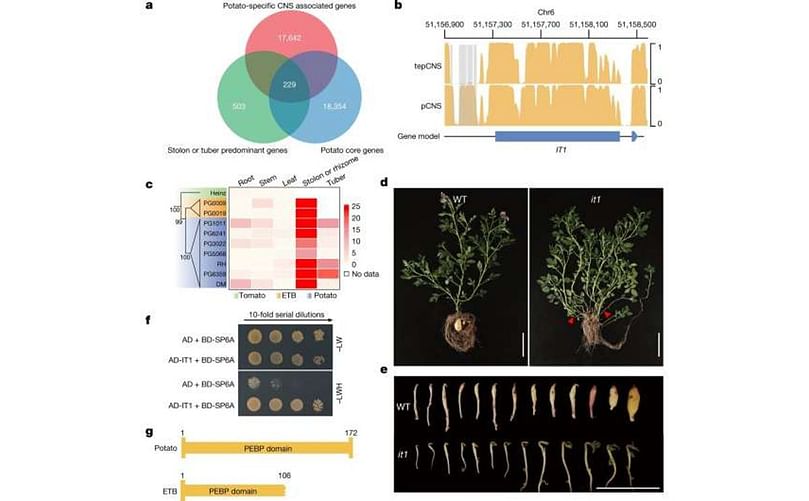
Identification of a potato tuber identity gene. a, Venn diagram describing the identification of 229 candidate
Glenn Bryan, a potato geneticist at the James Hutton Institute’s Cell and Molecular Sciences group and a co-author of the study:
"This work really gives a 'pan genome' view across a diverse set of diploid potatoes that span the domestication timeline of this important global food crop. This work will be a fantastic source of information for potato geneticists and breeders and will provide a very rich source of data that will facilitate diploid hybrid breeding."They suggest this is due to their tuber-based propagation characteristics and the ways that the varieties have adapted to different environments as they were exported around the globe by human farmers. They also found that potatoes are most closely related to tomatoes.
The researchers also used CRISPR–Cas9 gene editing on some varieties to learn more about tuber development. They discovered a previously unknown gene that plays a role in the development of tubers, while also realizing that the protein it signals for interacts with an important tuber development molecule.
The researchers conclude that work like theirs is becoming more important as farmers around the world will be forced to change established practices to continue providing the food that will be needed for a growing population.
Publication
Dié Tang et al, Genome evolution and diversity of wild and cultivated potatoes, Nature (2022). DOI: 10.1038/s41586-022-04822-x
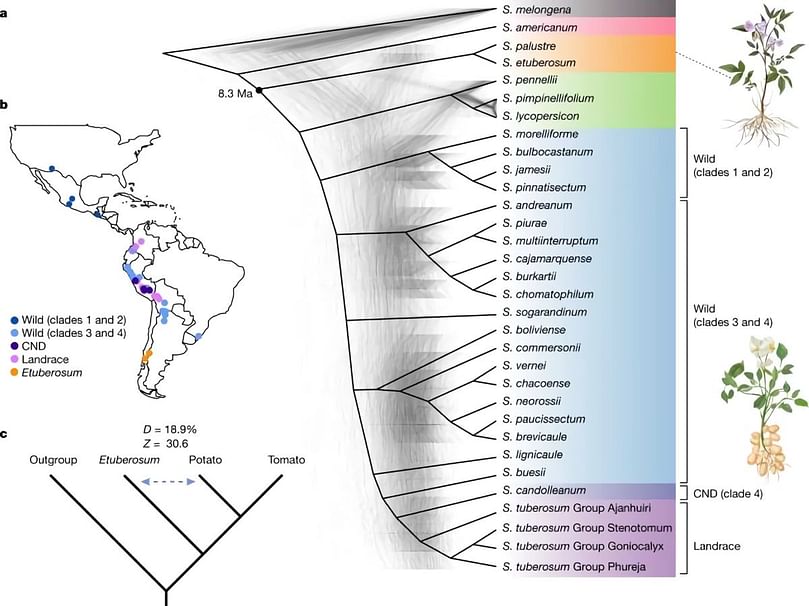
Geographical distribution and phylogeny of the Solanum genus.