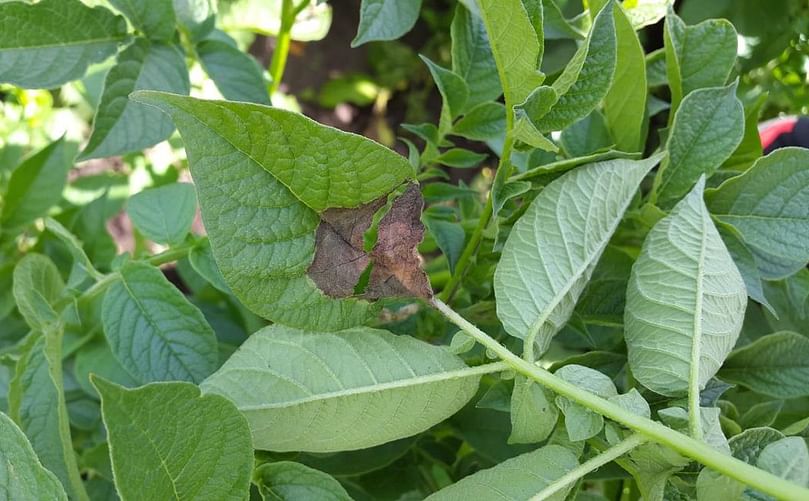
Mapping of the Phytophthora infestans genotypes found accross Europe in the EuroBlight sampling during 2017. In the article a link is provided to an interactive version of this map. (Courtesy: Euroblight)
Potato Blight Trends in Europe: EuroBlight 2017 Results
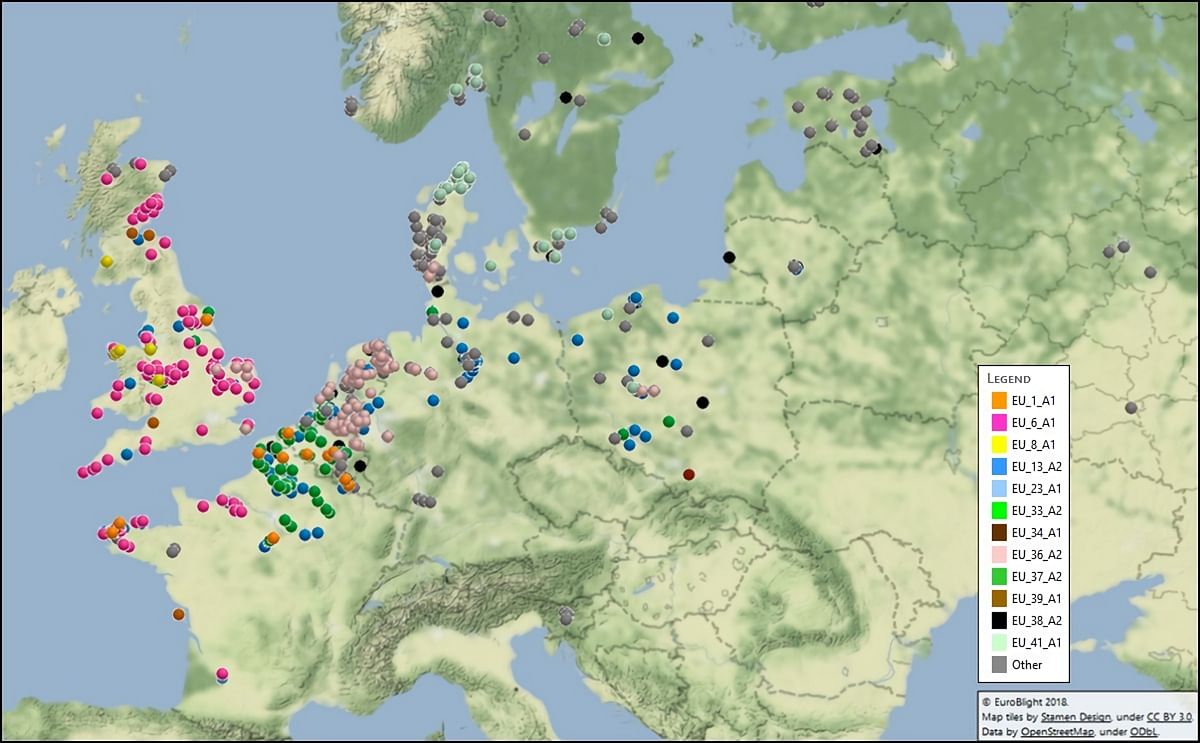
The EuroBlight project is examining the ongoing evolution of the potato late blight pathogen (Phytophthora infestans).
In 2017, almost 1500 samples from 16 countries in Europe were genotyped.
The results indicate that new clones continue to spread.
Latest findings
Around 75% of the samples belonged to defined clonal lineages also observed in previous seasons.
Some clones are widespread and have been present in Europe for more than a decade, but three emerging clones (EU_37_A2, EU_36_A2 and EU_41) increased their combined frequency from 10% in 2016 to 28% of the population in 2017.
Over the same period, genotype 13_A2 decreased from over 30% to less than 20% of the population.
A quarter of the population, mainly from the Nordic part of Europe, comprised ephemeral, genetically diverse isolates consistent with oospore-borne inoculum. A clear regional pattern in the dominance of clones versus sexual recombinants was observed across Europe.
Some implications of these displacements and ongoing changes are discussed.
Methodology
Since its arrival in the nineteenth century, Phytophthora infestans, the cause of potato late blight, has remained a serious threat to European potato production. Although we are now better equipped to control the disease than in the past, evolving pathogen populations continue to challenge integrated management practices.
Rapid changes in P. infestans populations causing late blight in Europe, America and Asia, including the emergence of strains with increased aggressiveness or reduced fungicide sensitivity, have been observed. Indeed, the changes in P. infestans populations directly influence the development and deployment of resistant cultivars, the performance of disease warning systems and the efficacy of plant protection products.
Therefore, coordinated and continuous pathogen monitoring was proposed by the EuroBlight consortium at its meeting in 2013, and now implemented as an EU-wide monitoring activity, including all stakeholders.
We continue to monitor populations and characterise the invasive genotypes to help optimise IPM strategies, as required by EU Directive 2009/128/EC on the sustainable use of plant protection products.

Example of a single sample FTA Card. FTA Cards are commonly used to take and preserve DNA Samples.
As in previous years, ‘FTA cards’ were distributed to disease ‘scouts’ across the industry, who visited blight-infected crops. Diseased leaves were pressed onto the cards and returned to the laboratories where the pathogen DNA was fingerprinted at the James Hutton Institute and INRA Rennes. The DNA fingerprint data defines the clonal lineages of the pathogen, and is combined with geo-location data to plot the diversity across Europe. Disease pressure in 2017 was lower than that in 2016, which reduced the sampling frequency and distribution slightly.
Around 1500 samples were collected by more than 20 partners across 16 European countries. As in previous years, the AHDB Potatoes ‘Fight Against Blight’ campaign data from crops in Great Britain (GB) is included. Included also is data from partners in the IPMBlight2.0 project, which is also generating pathogen phenotype data to support IPM strategies. The combined data from 2013-2017 now comprise over 6347 samples from 34 countries. With the support of international groups, data was also entered for parts of Asia, South America and North Africa.
Findings
Over the last five years, 60-79% of the population comprised known clonal lineages that recur each season. The remaining samples were novel, genetically diverse genotypes mostly found at a single location in one season and grouped in a category termed ‘Other’ (below).
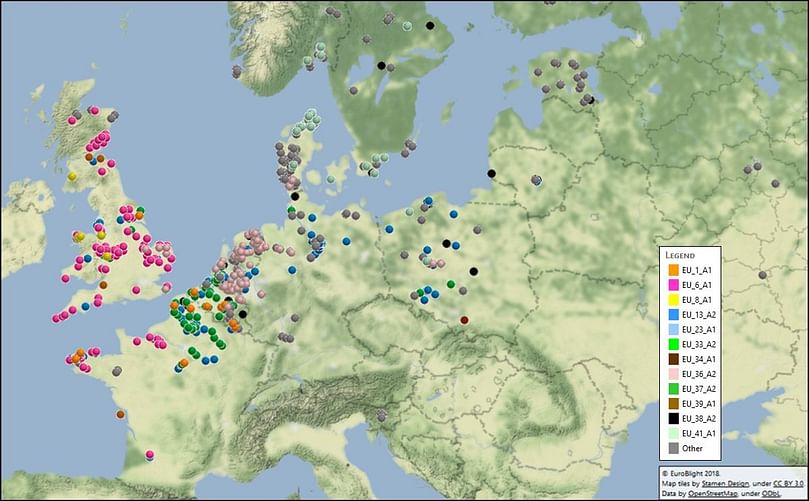
Mapping of the Phytophthora infestans genotypes found accross Europe in the EuroBlight sampling during 2017. Click image to access an interactive version of this map. (Courtesy: Euroblight 2018)
In 2017, clone EU_13_A2 (blue-13) declined from 31% of the samples to 19% which, for the first time, was lower than that of EU_6_A1 (23%).
The global distribution of the aggressive clone EU_13_A2 and its resistance to metalaxyl continues to affect management efficacy in Europe, parts of Asia and North Africa, reinforcing the need for pathogen data to support IPM best practices.
The frequency of EU_6_A1 was stable, but this clone remained localised to GB, France and a single sample from Belgium.
The frequency of EU_1_A1 further decreased from 4.5 to 2.2% of the population, and was found in the same countries as EU_6_A1.
Three new clones have been displacing the above genotypes and increased their range in 2017.
- Genotype EU_36_A2 was first sampled at low frequencies in the starch potato areas in Germany and the Netherlands in 2014 and 2015.
Over 2016 and 2017, its incidence increased from 3.7 to 8.8% of the population, and it has spread across the Netherlands into Belgium, GB, Denmark and Poland. - Genotype EU_37_A2 was first detected in Noordoostpolder in the Netherlands in 2013, and remained local at low frequency in 2014 and 2015.
It increased to 5.5% of the population sampled in 2016, and in 2017 reached 14.4%, also becoming established in England, Belgium and northern France. - Genotype EU_41 was first recorded in Denmark in 2013.
It has now become established, making up 4.5% of the 2017 population and having spread to southern Norway and Sweden and to northern Poland in a pattern consistent with airborne dispersal. The mating type test of EU_41 is pending.
The survival and spread of these clones, when others are decreasing or have failed to establish, suggests they are evolutionarily fit and may pose challenges to disease control.
Reduced sensitivity to fluazinam was reported in EU_37_A2 by Wageningen University in 2017, which is having implications on management practices. However, no information is yet available on the sensitivity of the other clones to fungicide active ingredients.
Lastly, the genetically diverse ‘Other’ samples comprised 25% of the population, and remained at the highest frequency in the east and north-east of Europe. In contrast to some clones, the traits possessed by the ‘Other’ isolates and other clones are not yet known. However, the IPMBlight2.0 project has started a detailed study of the traits of clonal and sexually recombinant isolates to help improve late blight management and forecasting.
The genetic diversity of the 2017 population can be visualised using an analysis tool (poppr 2.0) linked to the EuroBlight pathogen database. The minimum spanning network (below) shows the 2017 population structure, only including identified clones. Considerable sub-clonal diversity within the 13 year old EU_13_A2 lineage is observed compared to that in the ‘younger’ clones. ‘Other’ isolates (not shown) are genetically diverse and distributed across the whole network. Detailed analysis is underway to examine population change using these tools.
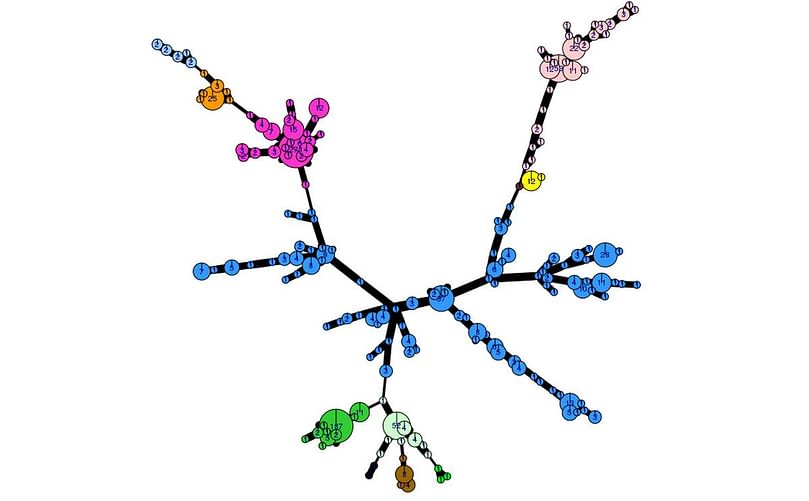
The genetic diversity of the 2017 Phytophthora infestans population visualised with poppr 2.0 (minimum spanning network)
The EuroBlight model of pathogen tracking is a rapid, cost-effective and co-ordinated approach to understanding pathogen evolution on a European scale. Data on the dominant clones has been passed to growers, advisors, breeders and agrochemical companies to provide practical management advice and shape longer-term strategies. The data provides an early warning of the incidence and spread of novel clones that is enabling a timely response from the industry.
The EuroBlight network is continuing to harmonise methods with other networks in the Americas and Asia and encourages continued co-operation between groups involved in managing late blight to exploit the database and tools for improved awareness and blight management on a global scale.