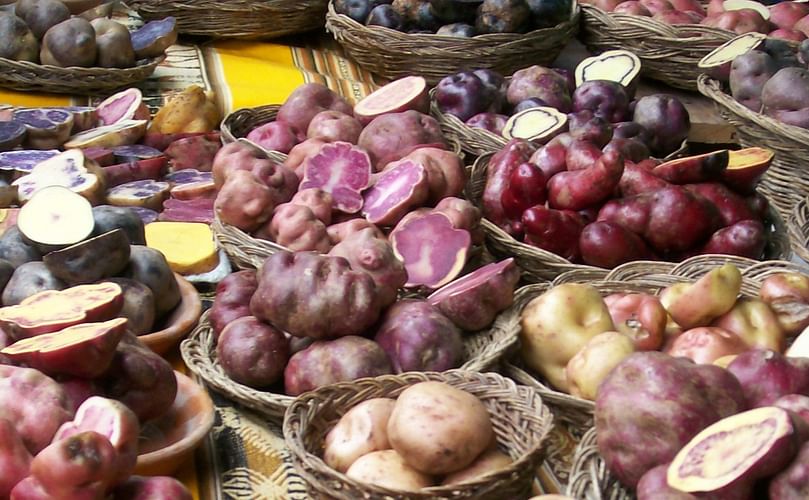
The enormous diversity of ancient / native potatoes can still be seen in countries like Peru, where many of these varieties are still grown.
Analyzing the genes of ancient potatoes helps to improve future potato varieties
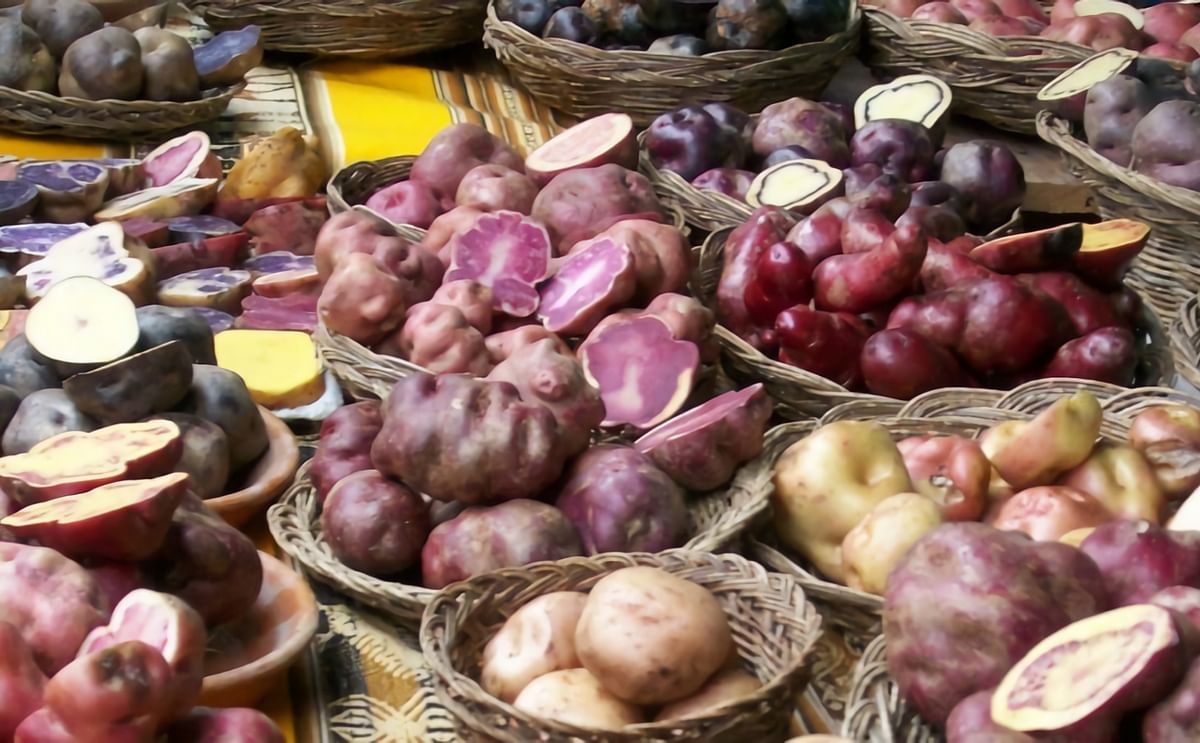

The old adage of looking to the past to understand the future certainly applies to improving potatoes.
Examining the ancestors of the modern, North American cultivated potato has revealed a set of common genes and important genetic pathways that have helped spuds adapt over thousands of years. The study appears in the current issue of Proceedings of the National Academy of Sciences.
Robin Buell, Michigan State University Foundation Professor of Plant Biology and senior author of the paper, shows potential genetic keys that could ensure the crop will thrive in the future.

Robin Buell, Michigan State University Foundation Professor of Plant Biology and senior author of the paper
Robin Buell:
“Worldwide, potato is the third most important crop grown for direct human consumption, yet breeders have struggled to produce new varieties that outperform those released over a century ago.”
“By analyzing cultivated potato and its wild relatives using modern genomics approaches, we were able to reveal key factors that could address food security in 21st century agriculture.”
Cultivated potatoes, domesticated from wild Solanum species, a genetically simpler diploid (containing two complete sets of chromosomes) species, can be traced to the Andes Mountains in Peru, South America.
While the exact means of the potato migration are unknown, spuds essentially spread worldwide since their domestication some 8,000 to 10,000 years ago. As potatoes were taken from the more equatorial regions of Peru and Bolivia to the southern parts of South America, they became adapted to longer summer days in Chile and Argentina.
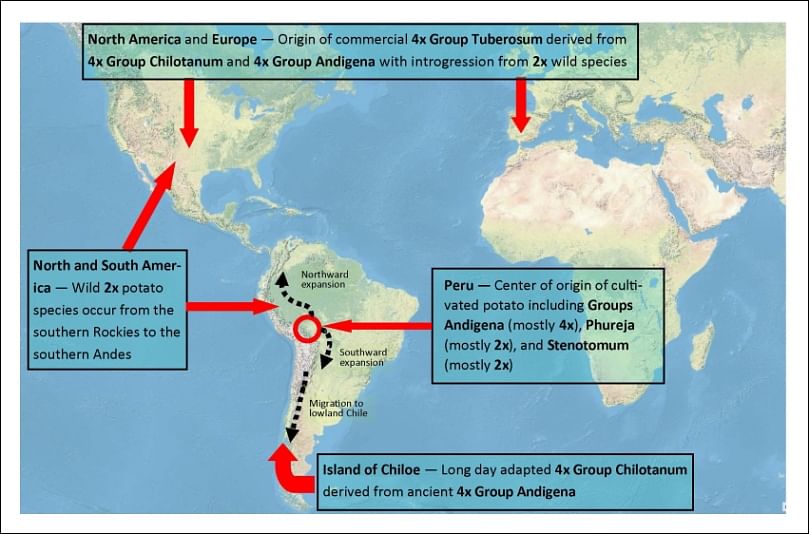
Potatoes spreading around the world - with some of their key genetic characteristics
One aspect that is known is how Spanish conquistadors introduced potatoes upon return from their South American exploits to the European continent, where potatoes were quickly adapted as a staple crop. As the explorers ventured from Europe to North America, they also brought potatoes to the new world.
Scientific explorer Michael Hardigan, formerly at MSU and now at the University of California-Davis, led the team of MSU and Virginia Polytechnic Institute and State University scientists. Together, they studied wild, landrace (South American potatoes that are grown by local farmers) and modern cultivars developed by plant breeders. The result, published in the current issue of Proceedings of the National Academy of Sciences, was the largest crop re-sequencing study to date.
Not only did it involve substantial re-sequencing of potato, but it also tackled one of the most-diverse crop genomes. The modern spuds found in today’s kitchens are genetically complex tetraploid potatoes, having four times the regular number of chromosomes. Potatoes’ complex genome harbors an estimated 39,000 genes. (In comparison, the human genome comprises roughly 20,000 genes.)
From the large gene pool, the researchers identified 2,622 genes that drove the crop’s early improvement when first domesticated. The study appears in the current issue of Proceedings of the National Academy of Sciences (open access).
Studying the gene diversity spectrum, from its wild past to its cultivated present, can provide an essential source of untapped adaptive potential, Buell said.
Robin Buell:
“We’ll be able to identify and study historic introgressions and hybridization events as well as find genes targeted during domestication that control variance for agricultural traits.”
“Many of these help focus on adapting to different climates, fending off different pathogens or improving yield, keys that we hope to better understand to improve future breeding efforts.”
For example, wild potatoes reproduce through berries and seeds. Cultivated potatoes are asexual and are food and seed in one. (Anyone who’s left a potato in a dark pantry too long has witnessed this trait firsthand.)
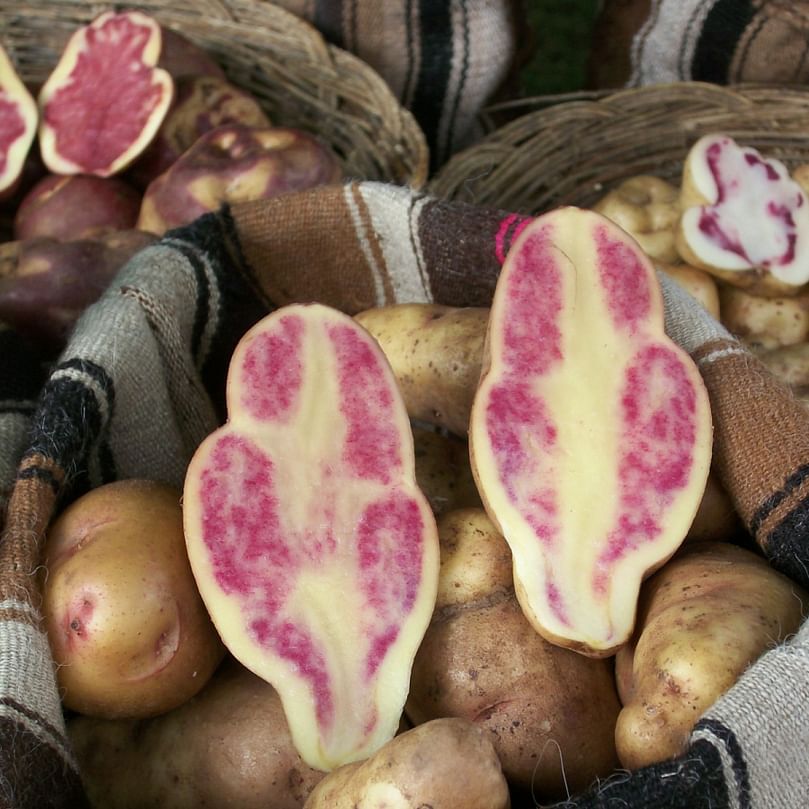

Studying the gene diversity spectrum, from its wild past to its cultivated present, can provide an essential source of untapped adaptive potential
The researchers present evidence of the signatures of selection in genes controlling this change. They also shed light on a role of wild species in genetic pathways for fighting pests and processing sugars for food. Diving into somewhat obscure territory, they looked at potential genetic sources that control circadian rhythm; yes, plants also have 24-hour clocks controlling biological processes.
Robin Buell:
“We knew about their physiological traits, but we didn’t know what genes were involved.”
“As potatoes were moved, they had to adapt to longer days, more hours of sunlight. We’re now starting to understand what’s happening at the genetic level and how wild Solanum species evolved to long-day adapted tetraploid potatoes.”
Additional MSU scientists contributing to this study include: Linsey Newton, John Hamilton, Brieanne Vaillancourt, Krystle Wiegert-Rininger, Joshua Wood, David Douches and Eva Farre. Scientists from Virginia Polytechnic Institute and State University also were part of the research team.